To the casual observer, Japanese stiltgrass appears as a harmless, leafy-green plant that blends into the majestic scenery during a weekend hike, but plant biologists like Craig Barrett know better.
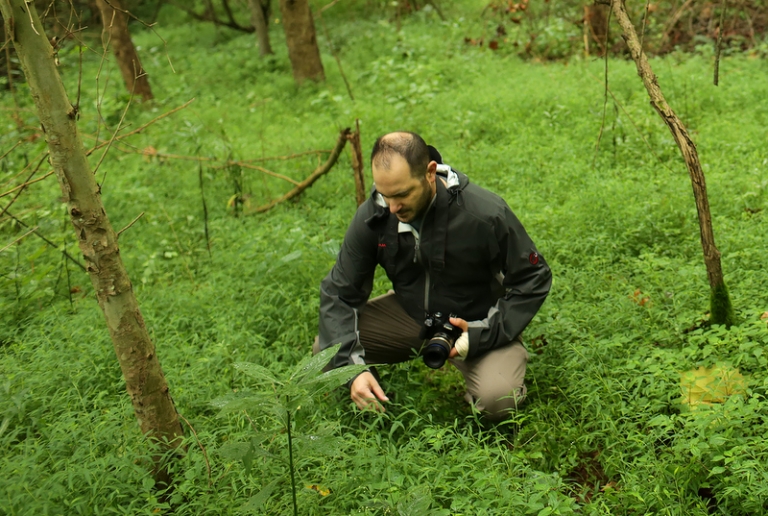
“I’d visited sites to study endangered orchids in West Virginia over the past few years, and I’d noticed Japanese stiltgrass creeping in and becoming a problem,” said Barrett, assistant professor of plant evolutionary biology at West Virginia University.

“So, in a way, you’re mad at this thing, and you want to study it—and there’s plenty of it to study.”
Japanese stiltgrass, or microstegium vimineum, is anything but harmless in the U.S.
It’s considered an invasive species, and it promotes disease, threatens biodiversity, negatively affects crops, restructures ecosystems, and damages infrastructure to the tune of $120 billion annually in the U.S.
It is believed that the stiltgrass was introduced into Tennessee around 1919, as it was used as packing material in shipments of porcelain from China.
It has since spread to at least 26 states across the East Coast, including West Virginia.
In Morgantown, West Virginia, it can easily be spotted at the Core Arboretum and along the municipal rail trail, Barrett said.
In hopes to ultimately ward-off or stabilize the grass—eradicating it is improbable at this point—the National Science Foundation has awarded Barrett and a team of researchers a highly competitive "Experimental Program to Stimulate Competitive Research" grant.
Barrett and his colleagues will receive $2 million to help them understand how plants undergo rapid evolution and become invasive and allow them to provide insights into the management and prevention of invasive species.
Barrett will sequence a complete genome of Japanese stiltgrass and collaborate with Cynthia Huebner, an adjunct professor at the Davis College of Agriculture, Natural Resources and Design, to conduct a greenhouse experiment using seeds from specimens collected in the U.S. and Asia.
The goal of Barrett’s project is to uncover the genomic basis of invasive traits using genome-wide sequencing methods and historical collections. Invasive species are still poorly represented among plants with completely sequenced genomes.
“Sequencing a genome is easy nowadays. The technology has changed so much in ten years,” Barrett said.
"We can easily go outside, collect plant material, grind it up in the lab, and send it to the sequencer. The hard part is interpreting the data. That takes the most time.”
One major element of a genome is called a transposable element—a DNA sequence that changes its position by “jumping around and affecting the functions of other genes,” he says. A transposable element can alter the cell’s genetic identity, drive changes in genome size, and change the way an organism interacts with its environment.
“The invasion of a new habitat, climate change, or getting attacked by a pathogen are events that can set off a cascade of activity of transposable elements in the genome,” Barrett explained.
“I’m interested in studying those responses both in the contemporary range and across time to see the changes in the abundance and diversity of the different transposable elements in the genome of this plant. Tracking this through time hasn’t really been done before.”
Results may indicate how rapid changes in the genome influence the spread of invasive species in new environments.
The project also involves scientists from Wichita State University, South Dakota State University, the University of Louisiana at Lafayette, and the University of Alabama at Tuscaloosa. A different high-profile invasive species will be studied at each of those universities.
“We have a handle on the ecology of invasion,” Barrett said.
“But figuring out how these species are adapting to new environments seems to be the question right now.
"I’m interested in how plants change over time both in terms of their genomes and physical features and how environmental factors might influence these changes.”